Velocity model to polygon simulPS (nnl)#
based on Daniel Trugman, 2024 + tips from C. Moody’s analysis notebook
Loïc Bachelot
import numpy as np
import pandas as pd
import pygmt
from pyproj import CRS, Transformer
import matplotlib.pyplot as plt
from scipy import interpolate
import xarray as xr
import os
import json
read json for all polygons to get the input parameters#
Poly id: int
olat, olon: origin for pyproj projection (center point)
plat1, plat2: lcc only (min and max of lat)
xmin, xmax, xsep = -124.0, 124.0, 2.0
ymin, ymax, ysep = -124.0, 124.0, 2.0
zmin, zmax, zsep = -4.0, 100.0, 1.0
cartesian grid bounds for velocity model - must encompass all stations, default values
with open("cascadia_poly.json", 'r+') as file:
file_data = json.load(file)
file_data[0]
{'poly_id': 1.0,
'olon': -123.7,
'olat': 48.0,
'plat1': 46.9,
'plat2': 49.1,
'xmin': -124.0,
'xmax': 124.0,
'xsep': 2.0,
'ymin': -124.0,
'ymax': 124.0,
'ysep': 2.0,
'zmin': -4.0,
'zmax': 100.0,
'zsep': 1.0,
'minlon': -125.0,
'maxlon': -122.4,
'minlat': 46.9,
'maxlat': 49.1}
setup parameter for the selected polygon#
### Setup
## Polygon Number: 1-10 and projection
poly = 3
pproj = "lcc" # "tmerc"
maxR = "200R"
print("Polygon", poly)
poly_param = file_data[poly-1]
# origin for pyproj projection
olat, olon = poly_param['olat'], poly_param['olon']
# lcc only
plat1, plat2 = poly_param['plat1'], poly_param['plat2']
# cartesian grid bounds for velocity model
# - must encompass all stations!
xmin, xmax, xsep = poly_param['xmin'], poly_param['xmax'], poly_param['xsep']
ymin, ymax, ysep = poly_param['ymin'], poly_param['ymax'], poly_param['ysep']
zmin, zmax, zsep = poly_param['zmin'], poly_param['zmax'], poly_param['zsep']
Polygon 3
set output path#
# output files
out_dir = "../locations/nlloc3D"
out_fileP = f"{out_dir}/model/delph_simulps_poly{poly}_{pproj}_{maxR}_P.txt"
out_fileS = f"{out_dir}/model/delph_simulps_poly{poly}_{pproj}_{maxR}_S.txt"
Define Projection#
# projection setup
crs1 = CRS.from_proj4("+proj=longlat +ellps=WGS84")
if pproj == "lcc":
crs2 = CRS.from_proj4("+proj={:} +ellps=WGS84 +lat_0={:.4f} +lon_0={:.4f} +lat_1={:.4f} +lon_2={:.4f} +units=km".format(
pproj,olat,olon,plat1,plat2))
else:
crs2 = CRS.from_proj4("+proj={:} +ellps=WGS84 +lat_0={:.4f} +lon_0={:.4f} +units=km".format(
pproj,olat,olon))
proj = Transformer.from_crs(crs1,crs2)
iproj = Transformer.from_crs(crs2,crs1)
Filter datasets and project#
load event list#
evlist = "../locations/cascadia.earthquakes.filtered"
qdf = pd.read_csv(evlist, parse_dates=['DateTime'])
# filter events for polygon
qdf = qdf[
(qdf['Longitude'] >= poly_param['minlon']) &
(qdf['Longitude'] <= poly_param['maxlon']) &
(qdf['Latitude'] >= poly_param['minlat']) &
(qdf['Latitude'] <= poly_param['maxlat'])
]
# event bounds
qlon0, qlon1 = qdf["Longitude"].quantile([0,1]).values
qlat0, qlat1 = qdf["Latitude"].quantile([0,1]).values
qlonM, qlatM = (qlon0+qlon1)/2, (qlat0+qlat1)/2
print(qlonM,qlatM)
# projection
qxp, qyp = proj.transform(qdf["Longitude"],qdf["Latitude"])
qdf["X"] = qxp
qdf["Y"] = qyp
# useful for NonLinLoc
print("Projected seismicity bounds:")
print(qdf["X"].min(), qdf["X"].max())
print(qdf["Y"].min(), qdf["Y"].max())
# show results
qdf
-123.69200000000001 46.0004
Projected seismicity bounds:
-98.34504694939974 102.3539043696198
-122.15082088039674 122.8226723669208
| DateTime | Latitude | Longitude | Depth | Magnitude | MagType | EventID | Source | X | Y | |
|---|---|---|---|---|---|---|---|---|---|---|
| 39 | 1970/09/03 03:38:37 | 46.8973 | -123.0958 | 46.9 | 2.1 | d | 10836893 | PNSN | 46.068818 | 99.953108 |
| 118 | 1971/05/28 10:42:07 | 46.5903 | -122.4315 | 14.6 | 3.4 | d | 10839063 | PNSN | 97.250671 | 66.395792 |
| 129 | 1971/06/16 22:49:32 | 46.6082 | -122.5233 | 44.5 | 1.7 | d | 10839158 | PNSN | 90.184398 | 68.280276 |
| 185 | 1971/09/14 13:31:33 | 46.4772 | -122.4320 | 11.3 | 2.5 | d | 10839748 | PNSN | 97.408813 | 53.819187 |
| 234 | 1971/12/13 20:59:03 | 47.0362 | -123.4942 | 27.3 | 3.6 | d | 10852228 | PNSN | 15.652714 | 115.253177 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 194365 | 2024/06/22 15:05:31 | 46.5368 | -122.4070 | 24.3 | 0.3 | l | 62017716 | PNSN | 99.223600 | 60.476475 |
| 194374 | 2024/06/23 03:12:40 | 46.3553 | -122.4153 | 16.2 | 2.1 | l | 62017906 | PNSN | 98.906146 | 40.285945 |
| 194462 | 2024/06/28 04:35:24 | 46.3570 | -122.4080 | 15.1 | 0.7 | l | 62011417 | PNSN | 99.465100 | 40.483872 |
| 194514 | 2024/07/01 04:21:36 | 46.1208 | -122.4978 | 18.9 | 0.7 | l | 62020301 | PNSN | 92.941290 | 14.118270 |
| 194515 | 2024/07/01 04:50:36 | 46.1232 | -122.5025 | 18.2 | 1.1 | l | 62020306 | PNSN | 92.574026 | 14.379700 |
6025 rows × 10 columns
load stations#
stlist = "../choosing_stations/cascadia.stations.filtered"
sdf = pd.read_csv(stlist, sep='|', parse_dates=['StartTime', 'EndTime'])
sdf
| Network | Station | Latitude | Longitude | Elevation | Sitename | StartTime | EndTime | Operating_Time | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | BK | DANT | 40.294601 | -121.802101 | 967.0 | Paynes Creek, CA, USA | 2024-06-27 17:40:00 | 2030-12-31 23:59:59 | 2378.263877 |
| 1 | C8 | BCOV | 50.544200 | -126.842700 | 33.0 | Beaver Cove, BC, CA | 2020-03-31 00:00:00 | 2030-12-31 23:59:59 | 3927.999988 |
| 2 | C8 | BPCB | 48.923600 | -123.704500 | 31.0 | Bare Point, BC, CA | 2008-03-19 00:00:00 | 2014-06-10 23:59:59 | 2274.999988 |
| 3 | C8 | FHRB | 50.060400 | -127.115800 | 5.0 | Fair Harbour, BC, CA | 2009-03-12 00:00:00 | 2015-10-24 23:59:59 | 2417.999988 |
| 4 | C8 | MWAB | 49.741100 | -125.303200 | 1176.0 | Mount Washington, BC, CA | 2009-07-31 00:00:00 | 2030-12-31 23:59:59 | 7823.999988 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 800 | CN | VGZ | 48.413000 | -123.325000 | 67.0 | Victoria Gonzales, BC, CA | 1994-07-21 00:00:00 | 2030-12-31 23:59:59 | 13312.999988 |
| 801 | CN | WOSB | 50.161000 | -126.570000 | 961.0 | Woss, BC, CA | 1998-11-17 00:00:00 | 2030-12-31 23:59:59 | 11732.999988 |
| 802 | CN | WPB | 49.648000 | -123.209000 | 260.0 | Watts Point, BC, CA | 1996-09-04 00:00:00 | 2030-12-31 23:59:59 | 12536.999988 |
| 803 | CN | WSLR | 50.127000 | -122.921000 | 907.0 | Whistler, BC, CA | 2003-05-09 00:00:00 | 2030-12-31 23:59:59 | 10098.999988 |
| 804 | CN | YOUB | 48.901000 | -124.262000 | 771.0 | Youbou Lake Cowichan, BC, CA | 2003-02-28 00:00:00 | 2014-09-03 23:59:59 | 4205.999988 |
805 rows × 9 columns
# load all stations
stlist = "../choosing_stations/cascadia.stations.filtered"
sdf = pd.read_csv(stlist, sep='|', parse_dates=['StartTime', 'EndTime'])
# filter stations for polygon
sdf = sdf[
(sdf['Longitude'] >= poly_param['minlon']) &
(sdf['Longitude'] <= poly_param['maxlon']) &
(sdf['Latitude'] >= poly_param['minlat']) &
(sdf['Latitude'] <= poly_param['maxlat'])
]
# find bounds
slat0, slat1 = sdf["Latitude"].min(), sdf["Latitude"].max()
slon0, slon1 = sdf["Longitude"].min(), sdf["Longitude"].max()
selv0, selv1 = sdf["Elevation"].min(), sdf["Elevation"].max()
print(f"min lon: {slon0}, max lon: {slon1}")
print(f"min lat: {slat0}, max lat: {slat1}")
print(f"min elevation: {selv0}, max elevation: {selv1}")
print()
# center point
slonC, slatC = np.round(0.5*(slon0+slon1),4), np.round(0.5*(slat0+slat1),4)
print(f"Center point lon: {slonC}, lat: {slatC}")
print()
# project coordinates
sxp, syp = proj.transform(sdf["Longitude"], sdf["Latitude"])
sdf["X"] = sxp
sdf["Y"] = syp
print(f"X min: {sdf["X"].min()}, X max: {sdf["X"].max()}")
print(f"Y min: {sdf["Y"].min()}, Y max: {sdf["Y"].max()}")
print()
# check bounds
assert sdf["X"].min() >= xmin
assert sdf["X"].max() <= xmax
assert sdf["Y"].min() >= ymin
assert sdf["Y"].max() <= ymax
# show dataframe
sdf
min lon: -124.07024, max lon: -122.401932
min lat: 45.02753, max lat: 47.08075
min elevation: 5.4, max elevation: 1130.0
Center point lon: -123.2361, lat: 46.0541
X min: -28.27793470777001, X max: 100.44969306512377
Y min: -108.0895647308553, Y max: 120.22042010062447
| Network | Station | Latitude | Longitude | Elevation | Sitename | StartTime | EndTime | Operating_Time | X | Y | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 20 | CC | CC | 45.610920 | -122.496130 | 66.66 | Rainier Lahar Test Station | 2019-01-01 | 2030-12-31 23:59:59 | 4382.999988 | 93.911013 | -42.554342 |
| 37 | CC | HUT1 | 45.611260 | -122.496760 | 86.00 | HUT 1 | 2023-02-10 | 2030-12-31 23:59:59 | 2881.999988 | 93.861311 | -42.517283 |
| 54 | CC | PR00 | 45.610920 | -122.496130 | 66.66 | Rainier Lahar Test Station | 2019-01-01 | 2030-12-31 23:59:59 | 4382.999988 | 93.911013 | -42.554342 |
| 95 | GS | TAUL1 | 45.522769 | -123.054179 | 56.00 | Cornelius Elementary School, Cornelius, OR,USA | 2021-05-25 | 2030-12-31 23:59:59 | 3507.999988 | 50.458219 | -52.848057 |
| 96 | GS | TAUL2 | 45.518911 | -123.109937 | 58.00 | Forest Grove Fire Dept, Forest Grove, OR, USA | 2021-05-26 | 2030-12-31 23:59:59 | 3506.999988 | 46.105025 | -53.310022 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 525 | UW | TOUT | 46.303180 | -122.563060 | 821.50 | Weyerhaeuser Mt St Helens Tree Farm, Cowlitz C... | 2023-06-04 | 2030-12-31 23:59:59 | 2767.999988 | 87.612402 | 34.321467 |
| 546 | UW | WHGC | 46.655991 | -123.729866 | 5.40 | Willapa Harbor Golf Course, WA, USA | 2019-02-03 | 2030-12-31 23:59:59 | 4349.999988 | -2.287108 | 72.942066 |
| 550 | UW | WPO | 45.572830 | -122.790527 | 335.40 | West Portland, OR, USA | 1986-10-16 | 2030-12-31 23:59:59 | 16147.999988 | 70.994361 | -47.086798 |
| 555 | UW | YACT | 45.932500 | -122.419300 | 214.00 | Yacolt, WA, USA | 2005-05-10 | 2030-12-31 23:59:59 | 9366.999988 | 99.339867 | -6.720267 |
| 556 | UW | YELM | 46.940160 | -122.590810 | 102.90 | Yelm Schools Facilities and Maintenance, Yelm,... | 2021-11-15 | 2030-12-31 23:59:59 | 3333.999988 | 84.506109 | 105.126743 |
91 rows × 11 columns
### Project
# create evenly spaced grid
xgrid, ygrid = np.meshgrid(
np.arange(xmin,xmax+xsep/2,xsep),
np.arange(ymin,ymax+ysep/2,ysep))
# print
print(xgrid)
print()
print(ygrid)
print()
# project
lon_grid, lat_grid = iproj.transform(xgrid,ygrid)
# print
print(lon_grid)
print()
print(lat_grid)
[[-124. -122. -120. ... 120. 122. 124.]
[-124. -122. -120. ... 120. 122. 124.]
[-124. -122. -120. ... 120. 122. 124.]
...
[-124. -122. -120. ... 120. 122. 124.]
[-124. -122. -120. ... 120. 122. 124.]
[-124. -122. -120. ... 120. 122. 124.]]
[[-124. -124. -124. ... -124. -124. -124.]
[-122. -122. -122. ... -122. -122. -122.]
[-120. -120. -120. ... -120. -120. -120.]
...
[ 120. 120. 120. ... 120. 120. 120.]
[ 122. 122. 122. ... 122. 122. 122.]
[ 124. 124. 124. ... 124. 124. 124.]]
[[-125.26931988 -125.24401442 -125.21870867 ... -122.18129133
-122.15598558 -122.13068012]
[-125.26980934 -125.24449599 -125.21918235 ... -122.18081765
-122.15550401 -122.13019066]
[-125.2702991 -125.24497786 -125.21965632 ... -122.18034368
-122.15502214 -122.1297009 ]
...
[-125.33137539 -125.3050698 -125.27876387 ... -122.12123613
-122.0949302 -122.06862461]
[-125.33190432 -125.30559021 -125.27927575 ... -122.12072425
-122.09440979 -122.06809568]
[-125.3324336 -125.30611095 -125.27978797 ... -122.12021203
-122.09388905 -122.0675664 ]]
[[44.87357522 44.87392034 44.87425986 ... 44.87425986 44.87392034
44.87357522]
[44.89156888 44.89191411 44.89225373 ... 44.89225373 44.89191411
44.89156888]
[44.90956248 44.90990782 44.91024754 ... 44.91024754 44.90990782
44.90956248]
...
[47.06784496 47.06820333 47.06855588 ... 47.06855588 47.06820333
47.06784496]
[47.08581831 47.08617679 47.08652944 ... 47.08652944 47.08617679
47.08581831]
[47.10379138 47.10414997 47.10450273 ... 47.10450273 47.10414997
47.10379138]]
load velocity model#
### Load xarray dataset
f_nc = "../locations/velocity_interp_delph2018.nc"
# load
velocity = xr.open_dataset(f_nc)
# show fields
velocity
<xarray.Dataset> Size: 15MB
Dimensions: (lat: 68, lon: 65, depth: 105)
Coordinates:
* lat (lat) float64 544B 36.0 36.2 36.4 36.6 ... 48.8 49.0 49.2 49.4
* lon (lon) float64 520B -124.8 -124.6 -124.4 ... -112.4 -112.2 -112.0
* depth (depth) int64 840B -4 -3 -2 -1 0 1 2 3 ... 94 95 96 97 98 99 100
Data variables:
Vp (lat, lon, depth) float64 4MB ...
Vs (lat, lon, depth) float64 4MB ...
Vs_interp (depth, lat, lon) float64 4MB ...
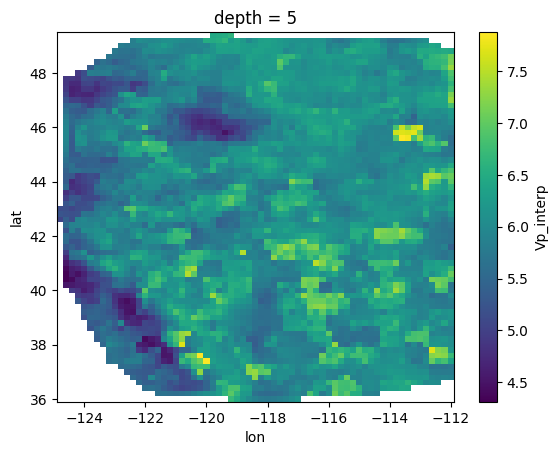
Vp_interp (depth, lat, lon) float64 4MB ...# fixed depth, map view
velocity['Vp_interp'].sel(depth = 5).plot()
plt.show()
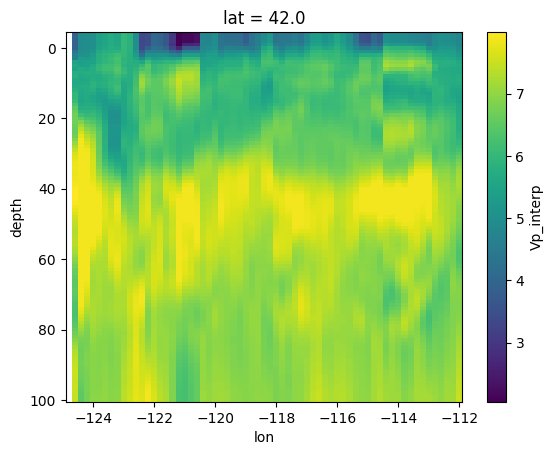
# cross section
velocity['Vp_interp'].sel(lat = 42).plot(y = 'depth')
plt.gca().invert_yaxis()
plt.show()
### Example PyGMT Map
# Depth for horizontal slice
depth = 0
# Setting variables related to the colorscale
vmin1 = 1.400
vmax1 = 5.200
vspace = 0.025
vs1_series = (vmin1, vmax1, vspace)
vs1_above = [vmax1,vmax1]
vs1_below = [vmin1,vmin1]
#Setting projection variables
projection = 'L-118/44/36/49/7.5c'
region = [-125,-112,36,50]
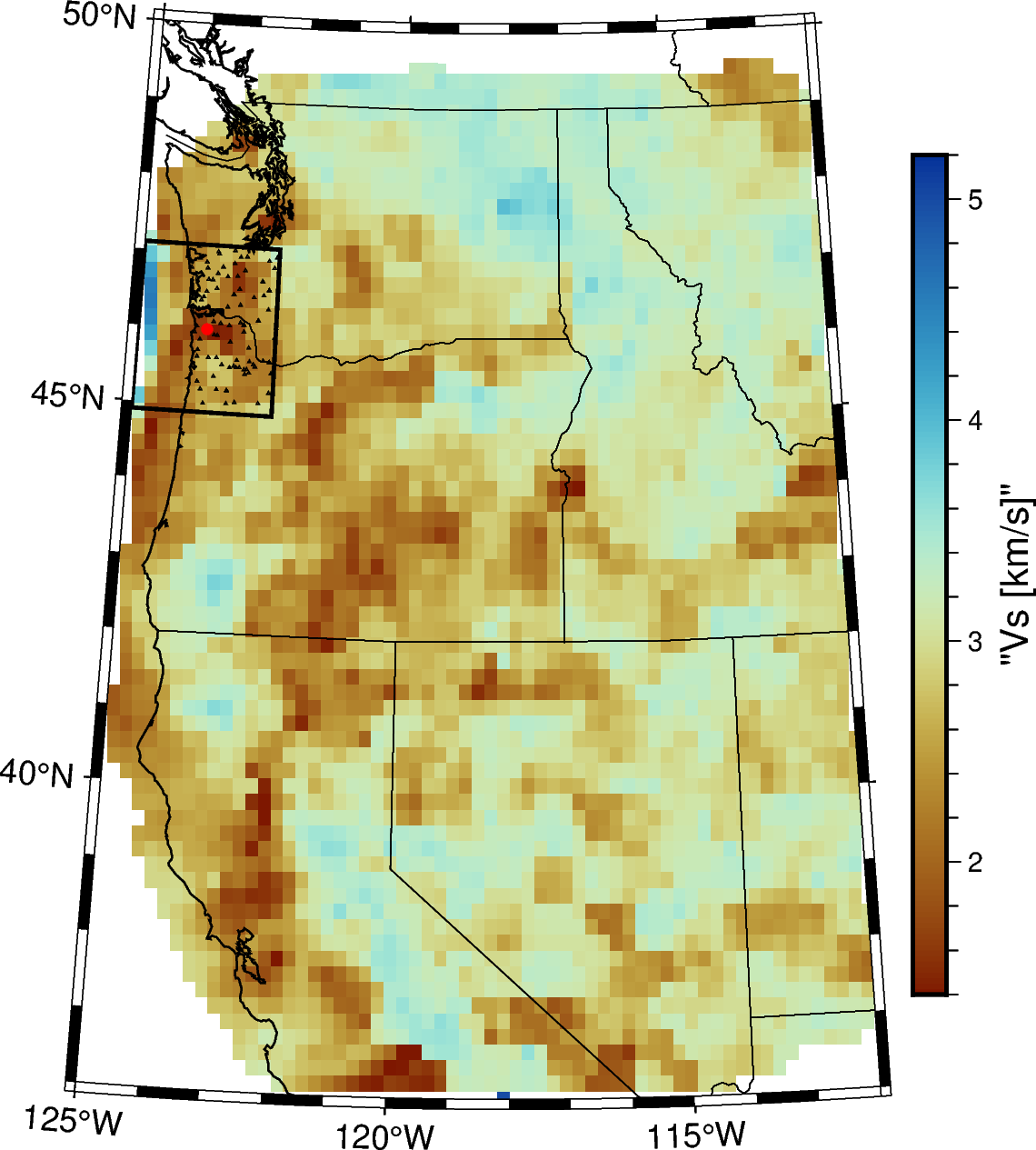
# map figure
fig = pygmt.Figure()
# make colormap
pygmt.makecpt(cmap="roma", series=vs1_series)
# Plot a 5km map view slice of the model in isotropic Vs
fig.grdimage(grid=pygmt.grdclip(velocity['Vs_interp'].sel(depth=depth),
below=vs1_below, above=vs1_above), projection=projection)
fig.colorbar(frame='af+l"Vs [km/s]"',position="JMR+o0.3c/0c")
# Coast
fig.coast(shorelines = '1/0.5p', region = region, projection=projection,
frame = ['af'], borders=["1/black", "2/black"])
# stations
fig.plot(x=sdf["Longitude"],y=sdf["Latitude"],style="t1p")
# origin
fig.plot(x=olon,y=olat,style="c3p",fill="red")
# selected polygon
fig.plot(x=[poly_param['minlon'], poly_param['minlon'], poly_param['maxlon'], poly_param['maxlon'], poly_param['minlon']],
y=[poly_param['minlat'], poly_param['maxlat'], poly_param['maxlat'], poly_param['minlat'], poly_param['minlat']]
, pen="1p, black")
# Show Results
# fig.savefig("./velocity_depth5.png", dpi=300)
fig.show()
project velocity to polygon#
### Compile DataFrame Using Scipy Interpolate
# - The Xarray version is much slower and crashes...
# first get flattened arrays for output grid
x_flat, y_flat = np.ravel(xgrid), np.ravel(ygrid)
out_lonflat, out_latflat = np.ravel(lon_grid), np.ravel(lat_grid)
# from velocity model
longrid, latgrid = np.meshgrid(
np.asarray(velocity.lon), np.asarray(velocity.lat))
print(longrid.shape)
vel_lonflat = np.ravel(longrid)
vel_latflat = np.ravel(latgrid)
# 2D arrays as input for scipy.interpolate
out_obs = np.vstack([out_lonflat, out_latflat]).T
vel_obs = np.vstack([vel_lonflat, vel_latflat]).T
# store data here
xvals, yvals, zvals, vpvals, vsvals = [], [], [], [], []
# loop over depths
for zz in np.arange(-4.0, zmax+zsep, zsep):
vel_vp = np.ravel(velocity['Vp_interp'].interp(depth = zz, method = 'linear').values)
vel_vs = np.ravel(velocity['Vs_interp'].interp(depth = zz, method = 'linear').values)
# interpolation
out_vp = interpolate.griddata(vel_obs, vel_vp, out_obs, method="linear")
out_vs = interpolate.griddata(vel_obs, vel_vs, out_obs, method="linear")
# update
xvals.append(x_flat)
yvals.append(y_flat)
zvals.append(zz + np.zeros(y_flat.size))
vpvals.append(out_vp) # km/s
vsvals.append(out_vs) # km/s
# progress
if np.mod(zz,10.0) < 0.1: print("Done with depth",zz)
# completed everything
print("Done")
(68, 65)
Done with depth 0.0
Done with depth 10.0
Done with depth 20.0
Done with depth 30.0
Done with depth 40.0
Done with depth 50.0
Done with depth 60.0
Done with depth 70.0
Done with depth 80.0
Done with depth 90.0
Done with depth 100.0
Done
## Turn This Into DataFrame
# make dataframe
odf = pd.DataFrame({
"x": np.hstack(xvals),
"y": np.hstack(yvals),
"z": np.hstack(zvals),
"vp": np.hstack(vpvals),
"vs": np.hstack(vsvals),
})
# grid points
xg= odf["x"].unique()
yg = odf["y"].unique()
zg = odf["z"].unique()
nx, ny, nz = len(xg), len(yg), len(zg)
print(nx, ny, nz)
# test groupby z and y
gdf = odf.groupby(["z","y"])
print(gdf.get_group((15.0,-100.0)))
# show
odf
125 125 105
x y z vp vs
298375 -124.0 -100.0 15.0 NaN NaN
298376 -122.0 -100.0 15.0 NaN NaN
298377 -120.0 -100.0 15.0 NaN NaN
298378 -118.0 -100.0 15.0 NaN NaN
298379 -116.0 -100.0 15.0 NaN NaN
... ... ... ... ... ...
298495 116.0 -100.0 15.0 6.192784 3.538732
298496 118.0 -100.0 15.0 6.195986 3.540562
298497 120.0 -100.0 15.0 6.193055 3.538888
298498 122.0 -100.0 15.0 6.189876 3.537071
298499 124.0 -100.0 15.0 6.186695 3.535254
[125 rows x 5 columns]
| x | y | z | vp | vs | |
|---|---|---|---|---|---|
| 0 | -124.0 | -124.0 | -4.0 | NaN | NaN |
| 1 | -122.0 | -124.0 | -4.0 | NaN | NaN |
| 2 | -120.0 | -124.0 | -4.0 | NaN | NaN |
| 3 | -118.0 | -124.0 | -4.0 | NaN | NaN |
| 4 | -116.0 | -124.0 | -4.0 | NaN | NaN |
| ... | ... | ... | ... | ... | ... |
| 1640620 | 116.0 | 124.0 | 100.0 | 7.325150 | 4.185799 |
| 1640621 | 118.0 | 124.0 | 100.0 | 7.362251 | 4.206999 |
| 1640622 | 120.0 | 124.0 | 100.0 | 7.399359 | 4.228203 |
| 1640623 | 122.0 | 124.0 | 100.0 | 7.434268 | 4.248152 |
| 1640624 | 124.0 | 124.0 | 100.0 | 7.441389 | 4.252221 |
1640625 rows × 5 columns
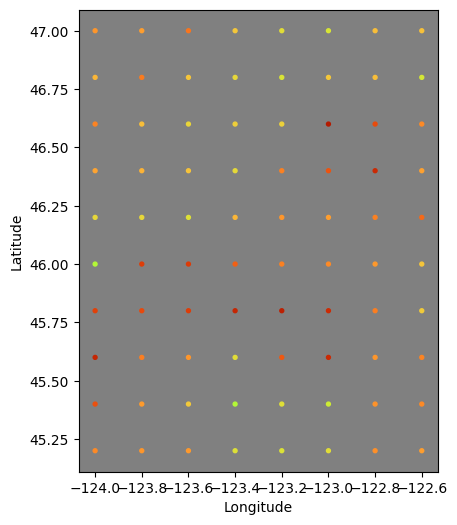
### Test Plot - Notice the Shallow NaN values for interp
test_depth = 0.0
# select data from the xarray
data = velocity["Vs_interp"].interp(depth = test_depth, method = "linear")
vmin = data.min().data
vmax = data.max().data
print(data)
print(f"vmin={vmin}, vmax={vmax}")
lons = np.asarray(data.lon)
lats = np.asarray(data.lat)
mx, my = np.meshgrid(lons,lats)
fx, fy = np.ravel(mx), np.ravel(my)
md = np.ravel(data.values)
# only within the study region bounds
idx = (fx>=slon0)&(fx<=slon1)&(fy>=slat0)&(fy<=slat1)
# plot results to check
fig, axi = plt.subplots(figsize=(6,6))
axi.set_aspect("equal")
axi.set_facecolor("gray")
axi.scatter(fx[idx],fy[idx],s=8,c=md[idx],vmin=vmin,vmax=vmax,
cmap=plt.cm.turbo_r)
axi.set_xlabel("Longitude")
axi.set_ylabel("Latitude")
plt.show()
# select data from dataframe
zdf = odf.groupby("z").get_group(test_depth)
# now plot results to check
fig, axi = plt.subplots(figsize=(6,6))
axi.set_aspect("equal")
axi.set_facecolor("gray")
axi.scatter(zdf["x"],zdf["y"],s=0.15,c=zdf["vs"], vmin=vmin, vmax=vmax,
cmap=plt.cm.turbo_r)
axi.set_xlabel("X [km]")
axi.set_ylabel("Y [km]")
plt.show()
<xarray.DataArray 'Vs_interp' (lat: 68, lon: 65)> Size: 35kB
array([[ nan, nan, nan, ..., nan, nan, nan],
[ nan, nan, nan, ..., nan, nan, nan],
[ nan, nan, nan, ..., nan, nan, nan],
...,
[ nan, nan, nan, ..., 2.5081, nan, nan],
[ nan, nan, nan, ..., nan, nan, nan],
[ nan, nan, nan, ..., nan, nan, nan]])
Coordinates:
* lat (lat) float64 544B 36.0 36.2 36.4 36.6 36.8 ... 48.8 49.0 49.2 49.4
* lon (lon) float64 520B -124.8 -124.6 -124.4 ... -112.4 -112.2 -112.0
depth float64 8B 0.0
vmin=1.0172, vmax=5.138352709337095
fill NaNs#
# Function to fill NaN values using nearest neighbors
def fill_nan_with_nearest_neighbors(df, columns):
for z_val in df['z'].unique():
subset = df[df['z'] == z_val]
for col in columns:
mask = subset[col].notna()
points = subset[mask][['x', 'y']].values
values = subset[mask][col].values
grid = subset[['x', 'y']].values
filled_values = interpolate.griddata(points, values, grid, method='nearest')
df.loc[df['z'] == z_val, col] = filled_values
return df
# Fill NaN values for vp and vs
odf = fill_nan_with_nearest_neighbors(odf, ['vp', 'vs'])
odf
| x | y | z | vp | vs | |
|---|---|---|---|---|---|
| 0 | -124.0 | -124.0 | -4.0 | 5.967288 | 3.409882 |
| 1 | -122.0 | -124.0 | -4.0 | 5.967288 | 3.409882 |
| 2 | -120.0 | -124.0 | -4.0 | 5.967288 | 3.409882 |
| 3 | -118.0 | -124.0 | -4.0 | 5.967288 | 3.409882 |
| 4 | -116.0 | -124.0 | -4.0 | 5.967288 | 3.409882 |
| ... | ... | ... | ... | ... | ... |
| 1640620 | 116.0 | 124.0 | 100.0 | 7.325150 | 4.185799 |
| 1640621 | 118.0 | 124.0 | 100.0 | 7.362251 | 4.206999 |
| 1640622 | 120.0 | 124.0 | 100.0 | 7.399359 | 4.228203 |
| 1640623 | 122.0 | 124.0 | 100.0 | 7.434268 | 4.248152 |
| 1640624 | 124.0 | 124.0 | 100.0 | 7.441389 | 4.252221 |
1640625 rows × 5 columns
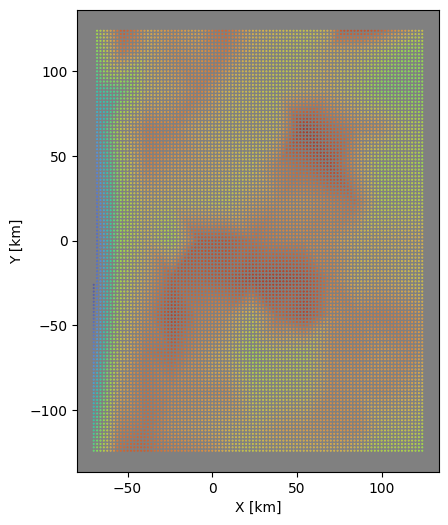
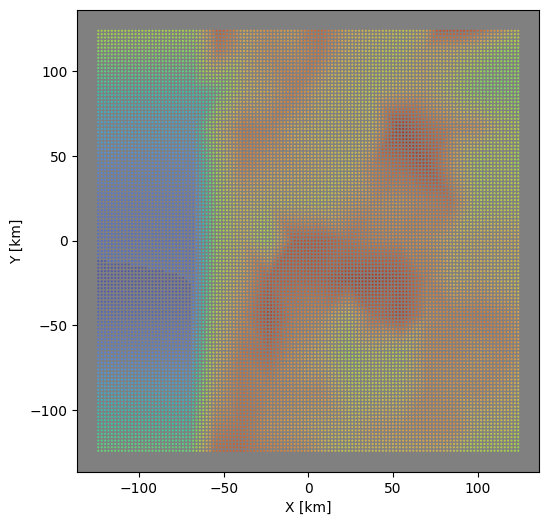
# select data from dataframe
zdf = odf.groupby("z").get_group(test_depth)
# now plot results to check
fig, axi = plt.subplots(figsize=(6,6))
axi.set_aspect("equal")
axi.set_facecolor("gray")
axi.scatter(zdf["x"],zdf["y"],s=0.15,c=zdf["vs"], vmin=vmin, vmax=vmax,
cmap=plt.cm.turbo_r)
axi.set_xlabel("X [km]")
axi.set_ylabel("Y [km]")
plt.show()
Save velocity grid#
### Write Vp Grid
# open output file
with open(out_fileP,"w") as f:
# header line
unit, nph = 1.0, 2 # grid units are 1km, 2 phases
f.writelines("%f %d %d %d %d # SimulPS P-wave Model\n" %(unit,nx,ny,nz,nph))
# write xgrid, ygrid, zgrid
s = " ".join("%.1f" %x for x in xg) + "\n"
f.writelines(s)
s = " ".join("%.1f" %y for y in yg) + "\n"
f.writelines(s)
s = " ".join("%.1f" %z for z in zg) + "\n"
f.writelines(s)
# dummy lines
f.writelines("0 0 0\n\n")
# loop over z, then y, then write all x
gdf = odf.groupby(["z","y"])
for zval in zg:
for yval in yg:
df = gdf.get_group((zval,yval))
vdata = df["vp"].values
s = " ".join("%.2f" %v for v in vdata) + "\n"
f.writelines(s)
# write results
print("Done with",out_fileP)
Done with ../locations/nlloc3D/model/delph_simulps_poly3_lcc_200R_P.txt
### Write Vs Grid
# open output file
with open(out_fileS,"w") as f:
# header line
unit, nph = 1.0, 2 # grid units are 1km, 2 phases
f.writelines("%f %d %d %d %d # SimulPS S-wave Model\n" %(unit,nx,ny,nz,nph))
# write xgrid, ygrid, zgrid
s = " ".join("%.1f" %x for x in xg) + "\n"
f.writelines(s)
s = " ".join("%.1f" %y for y in yg) + "\n"
f.writelines(s)
s = " ".join("%.1f" %z for z in zg) + "\n"
f.writelines(s)
# dummy lines
f.writelines("0 0 0\n\n")
# loop over z, then y, then write all x
gdf = odf.groupby(["z","y"])
for zval in zg:
for yval in yg:
df = gdf.get_group((zval,yval))
vdata = df["vs"].values
s = " ".join("%.2f" %v for v in vdata) + "\n"
f.writelines(s)
# write results
print("Done with",out_fileS)
Done with ../locations/nlloc3D/model/delph_simulps_poly3_lcc_200R_S.txt
print("DONE, Polygon",poly)
DONE, Polygon 3